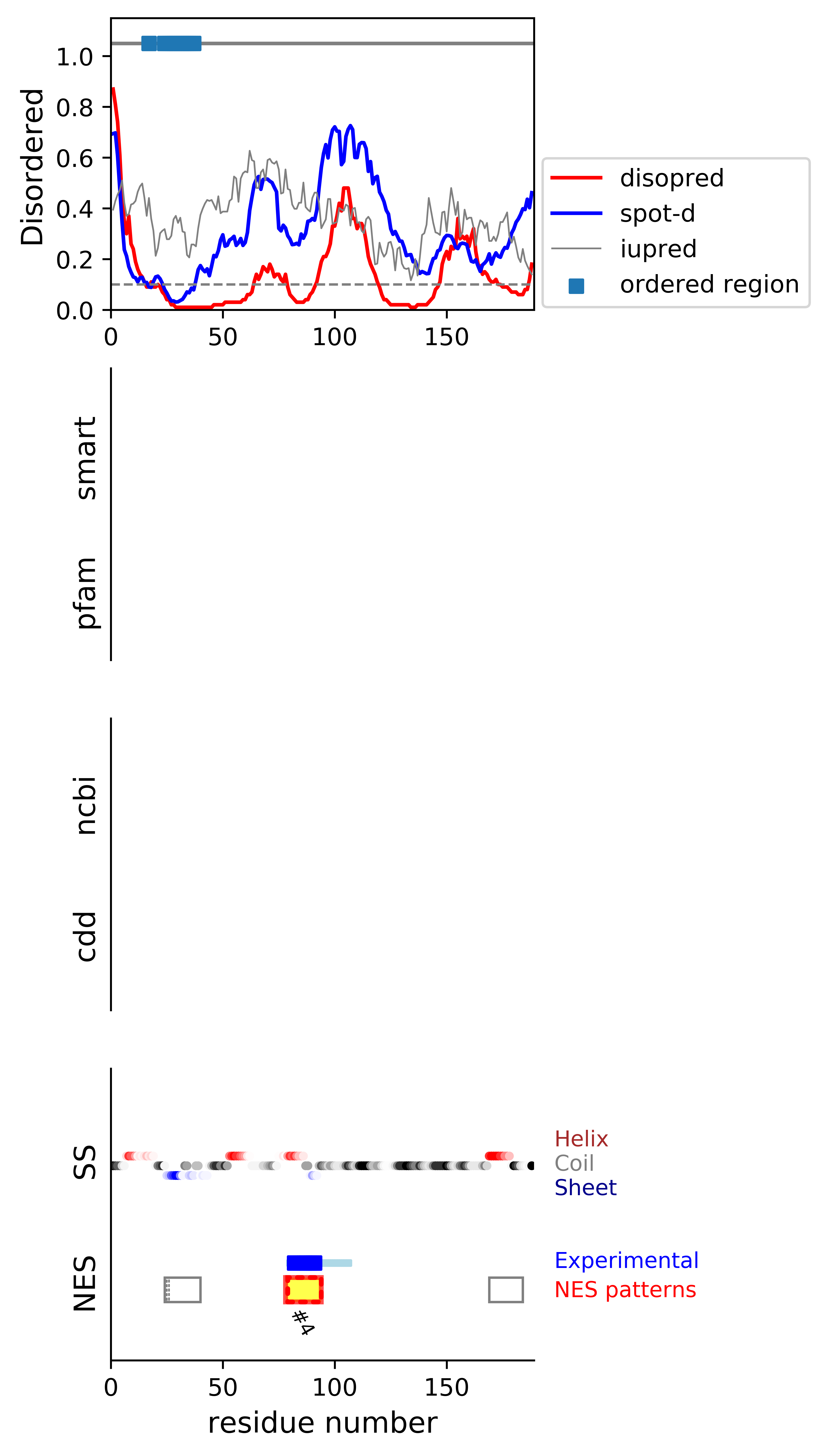

*Experimental: mutation (blue); functional region (cyan); located in the long functional sequence (lightblue)
*NES patterns: region with experimental evidence (red-orange-yellow); false positive region (gray)
| # | candidates | id | start# | sequence | secondary | class | multi-pattern | diso | spotd | iup | loc_DISO | loc_CDD | beta |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | fp_beta_D | Q83414 | 24 | EVFSFVFKTEDVQLNG | CEEEEEEEECCEEECC | c1a-AT-5 | multi-selected | 0.018 | 0.065 | 0.295 | boundary | 0.714 | |
| 2 | fp_beta_D | Q83414 | 25 | VFSFVFKTEDVQLNG | EEEEEEEECCEEECC | c1b-AT-5 | multi | 0.015 | 0.064 | 0.294 | boundary | 0.714 | |
| 3 | fp_beta_D | Q83414 | 26 | FSFVFKTEDVQLNG | EEEEEEECCEEECC | c2-AT-4 | multi | 0.014 | 0.064 | 0.295 | boundary | 0.714 | |
| 4 | cand_D | Q83414 | 78 | TVDEMTKKFGTLTIHD ..*+++**+***.. | HHHHHHHHHCCEEEEC | c1a-5 | multi-selected | 0.066 | 0.319 | 0.439 | DISO | 0.0 | |
| 5 | cand_D | Q83414 | 79 | VDEMTKKFGTLTIHD ..*+++**+***.. | HHHHHHHHCCEEEEC | c1b-AT-4 | multi | 0.061 | 0.318 | 0.432 | DISO | 0.143 | |
| 6 | fp_D | Q83414 | 169 | VEERLQRAAELGLRP | HHHHHHHHHHCCCCC | c1b-AT-4 | uniq | 0.089 | 0.263 | 0.289 | DISO | 0.0 |
*candidates: NES candidates and false positives annotated with "cand" and "fp", respectively;
if the segment is located in the disordered or boundary region, flagged with "D"; if the segment is located in the ordered region, flagged with "O";
if the segment's beta-strand content is over 0.5, flagged with "beta".
*sequence: Hydrophobic positions are colored in red. The positions with the experimental evidence is marked with '*' (mutation) and '+' (functional sequence in NESdb or sites in validNES).
The positions with '.' are for the region annotated in the long (more than 25 residues) functional sequence or site.
*multi-pattern: the consensus pattern is unique or multiple within the region (if the start# difference is less than 5, the segments are considered to be the same region)
*diso: average DISOPRED3-predicted disorder propensity for the segment
*spotd: average SPOT-Disorder-predicted disorder propensity for the segment
*iup: average IUPRED2A-predicted disorder propensity for the segment
*loc_DISO: location of the segment with respect to the ordered/disordered regions
*loc_CDD: location of the segment with respect to the conserved region annotated in the Conserved Domain Database (CDD)
*beta: beta-strand content in the middle of the segment