Rossmann-Like Proteins: Function and Evolution Analysis of a Fifth of the Protein World
The classical Rossmann fold, also known as a doubly-wound three layer a/b-sandwich, consists of two-units (321456 topology) that form a single parallel sheet flanked by alpha-helices on both sides and contain a characteristic crossover between strands 3 and 4. We defined its core minimal Rossmann-like motif (RLM) unit of three beta-strands flanked by two alpha-helices and found all known protein structures containing the RLM. We show that RLM enzymes function predominantly in metabolism, covering 38% of reference metabolic pathways. We find that closely related RLM enzyme families can catalyze different reaction chemistries using similar folds. Alternatively, different RLM folds can converge on catalyzing the same reactions. We showed that RLM enzymes utilize ligands from 20 chemical superclasses of organic and inorganic compounds. Homologous RLM domains can exhibit diverging active sites that accommodate alternate ligands, but with similar binding modes. The Rossmann fold is considered one of the most ancient folds, utilizing iron-sulfur clusters as cofactors and being the part of ancient energy metabolism, the Wood-Ljungdahl pathway, used by LUCA. Our data suggests that the top three disease categories with mutations in RLM proteins are diseases of endocrine system, nervous system and developmental anomalies.
Minimal Rossmann-like motif (RLM) definition
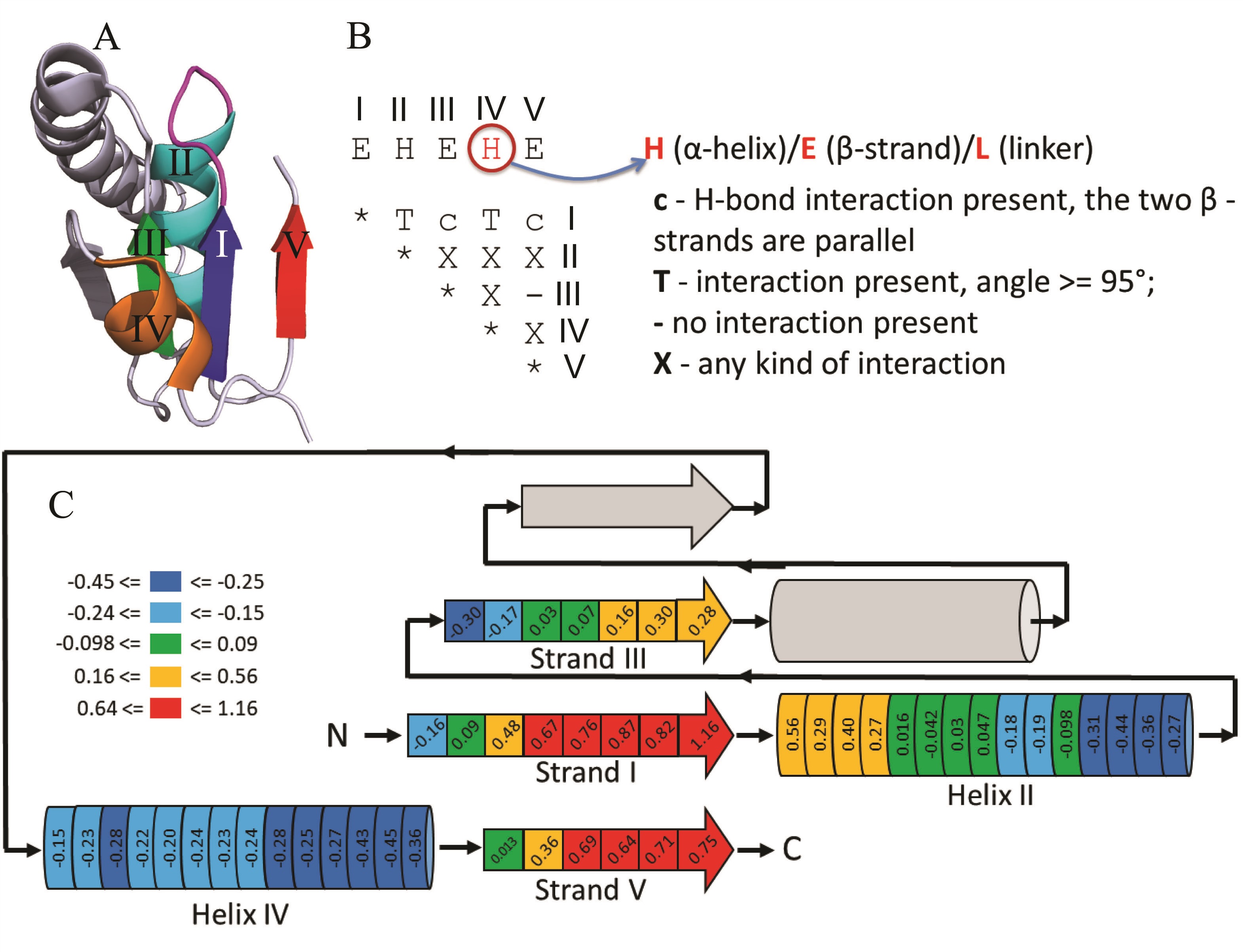
(A) RLM SSEs adapted from 5-formly-3-hydroxy-2-methylpyridine 4-carboxylic acid (FHMPC) 5-dehydrogenase (PDB: 4OM8) are numbered and colored in rainbow, with magenta catalytic loop between first β-strand - element I (β1) and first α-helix - element II (α1). The second α-helix - element IV (α2) forms crossover between second β-strand - element III (β2) and third β-strand - element V (β3). The crossover loop is unstructured loop at the N-terminal part of α2. Element IV can be α-helix, β-strand, or loop. The unlabeled SSEs (colored in slate) are considered as an insertion to the RLM, which can occur between element III (β2) and element IV (α2) or in any of the loops connecting the RLM SSEs.
(B) An interaction matrix defines RLM search strategy using ProSMoS program. Interaction type “T” considers the angle between vectors corresponding to particular RLM elements.
(C) RLM scheme with average AL2CO positional conservation index among family level representatives. RLM bins are colored according to conservation index from blue (not conserved) to red (highly conserved), non-RLM elements are shown in gray. Left side of all SSE corresponds to N-terminus, right side to the C-terminus.
Papers and Data:
1) Kirill E. Medvedev, Lisa N. Kinch, R. Dustin Schaeffer, Jimin Pei, Nick V. Grishin (2020) A Fifth of the Protein World: Rossmann-like Proteins as an Evolutionarily Successful Structural Unit. Journal of Molecular Biology 433(4): 166788 PMID: 33387532
Data based on ECOD version 20200517 (develop275):
a) Minimal Rossmann-like motif (RLM) containing homology groups
b) Conserved core of RLM-containing topology groups
c) Comparison of RLM-containing domains classification in new (v275) and old ECOD versions
d) Comparison of RLM-containing domains classification in ECOD and SCOP2
2) Kirill E. Medvedev, Lisa N. Kinch, R. Dustin Schaeffer, Nick V. Grishin (2019) Functional analysis of Rossmann-like domains reveals convergent evolution of topology and reaction pathways. Plos Computational Biology 15(12): e1007569 PMID: 31869345
Data based on ECOD version 20181017 (develop214):
a) Data set of RLM containing PDBs. Each PDB mapped to UniProt KB, ECOD database, EC number (if available). Substrate and Product information was retrieved from KEGG, Modified residues, Nucleic acid contact and Polysaccharide contact—from Protein Data Bank.
c) Classification of ligands associated with RLM EC numbers based on ClassyFire database
3) Kirill E. Medvedev, Lisa N. Kinch, Nick V. Grishin (2018) Functional and evolutionary analysis ofviral proteins containing a Rossmann-like fold. Protein Science 27(8): 1450-1463 PMID: 29722076
Data based on ECOD version 20161205 (develop159):
a) Viral protein families, which contain Rossmann-like domains