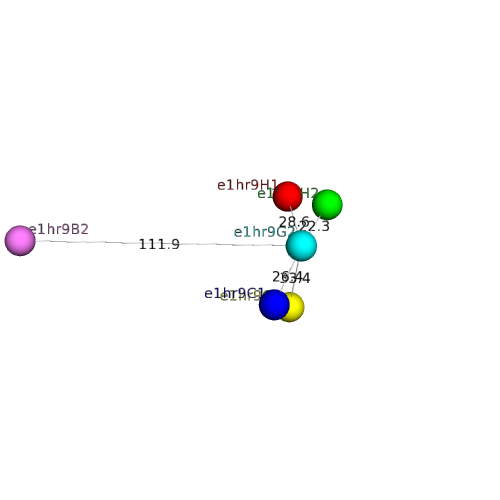
| Domain ID | Symmetry operator | H-group | Visualization |
|---|
| e1hr9H2 |
G:X,Y,Z->H:x,y,z |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|
| e1hr9H1 |
G:X,Y,Z->H:x,y,z |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|
| e1hr9C2 |
G:x,y,z->C:-X+1,Y-1/2,-Z+1/2 |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|
| e1hr9C1 |
G:x,y,z->C:-X+1,Y-1/2,-Z+1/2 |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|
| e1hr9B2 |
G:X-1,Y,Z->B:x,y,z |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|
| e1hr9E1 |
G:x,y,z->E:X,Y,Z |
LuxS, MPP, ThrRS/AlaRS common domain |
Interaction
Interface
Pymol
|