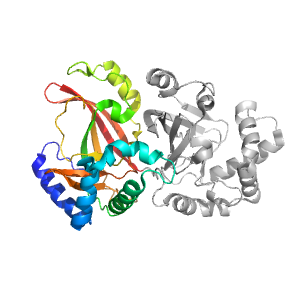
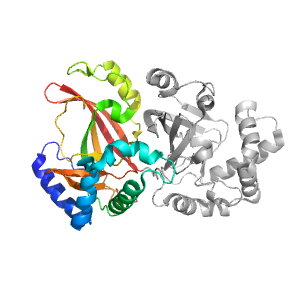
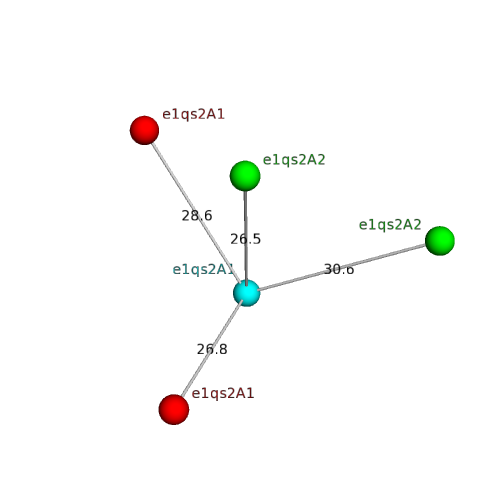
| Domain ID | Symmetry operator | H-group | Visualization |
|---|
| e1qs2A1 |
A:x,y,z->A:-X,Y-1/2,-Z+1/2 |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A1 |
A:-X,Y-1/2,-Z+1/2->A:x,y,z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A2 |
A:-X,Y-1/2,-Z+1/2->A:x,y,z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A2 |
A:x,y,z->A:X-1/2,-Y+1/2,-Z+1 |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A2 |
A:-X+1,Y-1/2,-Z+1/2->A:x,y,z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A2 |
A:X,Y-1,Z->A:x,y,z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A1 |
A:x,y,z->A:X-1,Y,Z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A2 |
A:x,y,z->A:X-1,Y,Z |
ADP-ribosylation |
Interaction
Interface
Pymol
|
| e1qs2A1 |
A:X-1,Y,Z->A:x,y,z |
ADP-ribosylation |
Interaction
Interface
Pymol
|