| UID: | 000436619 |
|---|---|
| Type: | Automatic domain |
| Parent: | e1x38A1 |
| Range: | A:5-396 |
| Ligands: | CA EDO K NA |
| PDB: | 3usz |
| PDB Description: | EXO-1,3/1,4-BETA-GLUCANASE |
| UniProt: | Q0QJA3 |
| Species: | Pseudoalteromonas sp. BB1 |
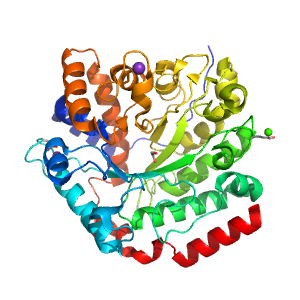
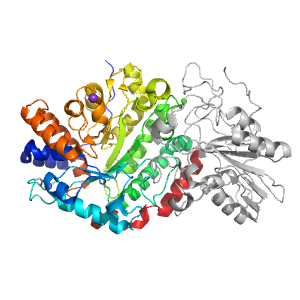
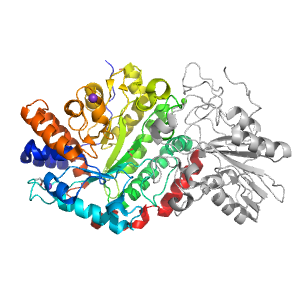
Structure of domain e3uszA1
Domains in the same chain:
| e3uszA1 | A:5-396 | TIM beta/alpha-barrel | TIM barrels | TIM barrels |
| e3uszA2 | A:391-626 | Flavodoxin-like | Beta-D-glucan exohydrolase, C-terminal domain | Beta-D-glucan exohydrolase, C-terminal domain |
Domains in the same PDB:
| e3uszA1 | A:5-396 | TIM beta/alpha-barrel | TIM barrels | TIM barrels |
| e3uszA2 | A:391-626 | Flavodoxin-like | Beta-D-glucan exohydrolase, C-terminal domain | Beta-D-glucan exohydrolase, C-terminal domain |
Structural contacts of domain e3uszA1 of H-group "TIM barrels":
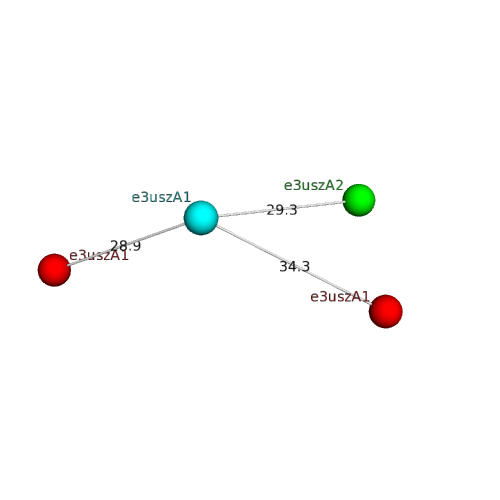
The domain and its neighboring domains (both within the asymmetric unit and to crystallographic symmetry mates) are represented by spheres and linked by lines. Distances between the center of the domain and interfaces are shown for each contact.
| Domain ID | Symmetry operator | H-group | Visualization |
|---|---|---|---|
| e3uszA1 | A:x,y,z->A:X,-Y,-Z | TIM barrels | Interaction Interface Pymol |
| e3uszA2 | A:x,y,z->A:X,-Y,-Z | Beta-D-glucan exohydrolase, C-terminal domain | Interaction Interface Pymol |
| e3uszA2 | A:X,-Y,-Z->A:x,y,z | Beta-D-glucan exohydrolase, C-terminal domain | Interaction Interface Pymol |
| e3uszA1 | A:X,-Y,-Z->A:x,y,z | TIM barrels | Interaction Interface Pymol |
| e3uszA1 | A:x,y,z->A:X-1/2,-Y+1/2,-Z | TIM barrels | Interaction Interface Pymol |
| e3uszA1 | A:X-1/2,-Y+1/2,-Z->A:x,y,z | TIM barrels | Interaction Interface Pymol |